Figures & data
Figure 1. Location map of 23 sampling sites (circles filled white), showing the town of San José de Ocoa (see a description of the rock/landform attributes of the sites in ). The overlapping points are depicted slightly displaced from their actual positions to ensure that all become visible. The background is a color shaded-relief view (red-white is highland, green is lowland) based on a 30-m SRTM DEM (Ref: NASA LP DAAC, 2000. SRTM 1 Arc-Second Global, https://earthexplorer.usgs.gov/. Published September 2014). The bottom-left inset shows the area in the context of the Dominican Republic
Figure 2. Dendrogram of groups of sites based on selected environmental variables. Groups are indicated by bold letters A to F, and sites are denoted by numbers. See text for details and for a description of the transects
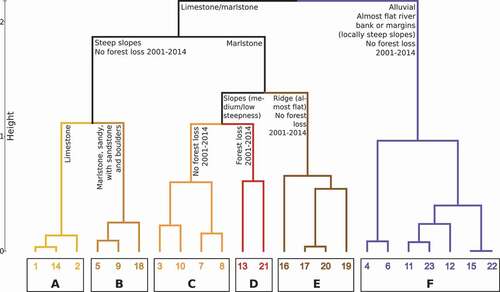
Table 1. IndVal calculations for 11 significantly associated species with groups of sites
Table 2. Rpb calculations for 13 significantly associated species with groups of sites
Table 3. List of species significantly associated with one or more groups of sites by means of IndVal and/or rpb.
Figure 3. Triplot of the Hellinger-transformed abundance community matrix, constrained by the variables (blue outlined rectangles) lithology, geomorphology, forest loss 2001–2014 and slope. Though we used the entire community matrix for computing the RDA, we only plotted the 16 species associated by means of IndVal and rpb. This is a scaling 2 triplot, where angles between species and explanatory variables, and angles between species themselves and variables themselves, are interpreted as correlations. The labelled (A to F) ellipses with grey dotted border enclose site groups defined by the AGNES agglomerative hierarchical clustering algorithm. See text for details and for a description of the transects
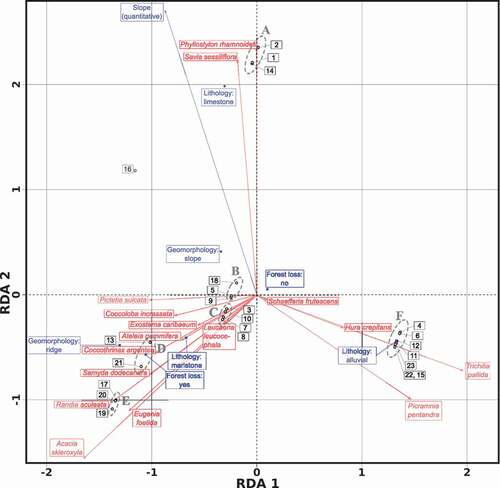
Table 4. Regression results from fitting the environmental variables onto a PCA ordination of a Hellinger transformed community matrix
Table A1. Morpho-topographic, geological and land use attributes of the transects
