Figures & data
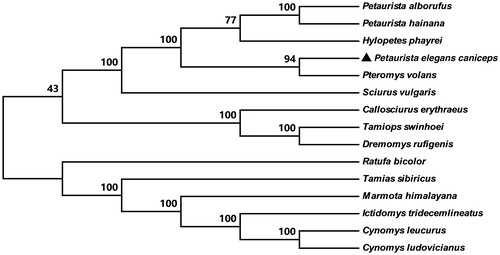
Figure 1. NJ tree inferred from the nucleotide sequence of whole mitogenome. The numbers above branches specify bootsrap percentages (1000 replicates). The species were selected from Petaurista elegans caniceps; Pteromys volans (JQ230001); Hylopetes phayrei (KC447305); Petaurista alborufus (JQ743657); Petaurista hainana (JX572159); Sciurus vulgaris (AJ238588); Dremomys rufigenis (KC447304); Tamiops swinhoei (KP027416); Tamias sibiricus (KF668525); Callosciurus erythraeus (KM502568); Ictidomys tridecemlineatus (KP698974); Cynomys leucurus (KP326309); Cynomys ludovicianus (KP326310); Marmota himalayana (JX069958); Ratufa bicolor (KF575124).