Figures & data
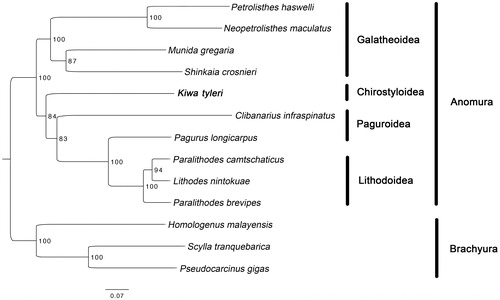
Figure 1. Maximum likelihood (ML) tree based on Kiwa tyleri with other 9 species from Anomura and 3 species from Brachyura. Bootstrap support values were generated with a rapid bootstrapping algorithm for 1000 replicates. The following mitogenomes were used in this analysis: Petrolisthes haswelli (LN624374), Neopetrolisthes maculatus (NC_020024), Munida gregaria (KU521508), Shinkaia crosnieri (EU420129), Clibanarius infraspinatus (LN626968), Pagurus longicarpus (AF150756), Paralithodes camtschaticus (JX944381), Lithodes nintokuae (AB769476), Paralithodes brevipes (AB735677), Homologenus malayensis (NC_026080), Scylla tranquebarica (NC_012567) and Pseudocarcinus gigas (NC_006891).