Figures & data
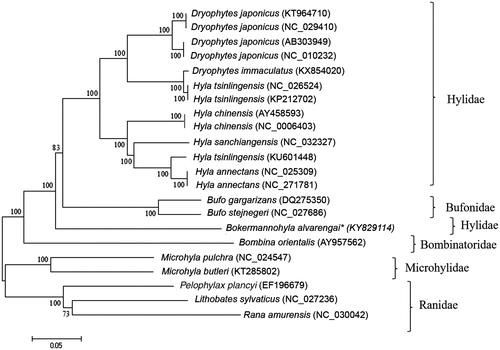
Figure 1. Phylogenetic tree generated using neighbor-joining algorithm based on complete mitochondrial genomes of the anuran species Dryophytes japonicus, D. immaculatus, Hyla tsinlingensis, H. chinensis, H. sanchiangensis, H. annectans, Bokermannohyla alvarengai, Bufo gargarizans, B. stejnegeri, Bombina orientalis, Microhyla pulchra, M. butleri, Pelophylax plancyi, Lithobates sylvaticus, and Rana amurensis. The phylogenetic tree was constructed under the Kimura-2 parameter model and consensus tree using 1000 bootstrap. Numbers indicate support of each clade.