Figures & data
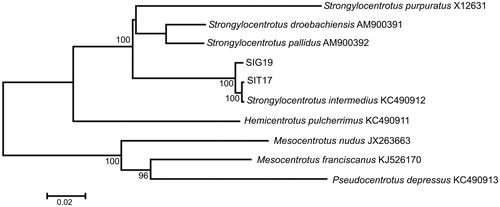
Figure 1. Maximum likelihood tree for the short-spined sea urchin Strongylocentrotus intermedius specimens SIG19 and SIT17, and GenBank representatives of the class Echinoidea. The tree is constructed using whole mitochondrial genomes. The tree is based on the general time reversible + gamma + invariant sites (GTR + G + I) model of nucleotide substitution. The numbers at the nodes are bootstrap percent probability values based on 500 replications (values below 75% are omitted).