Figures & data
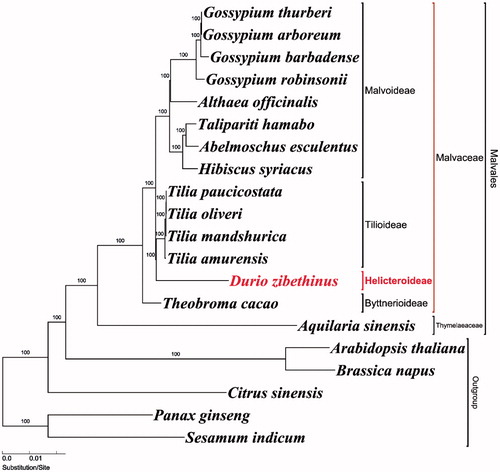
Figure 1. Maximum-likelihood (ML) tree based on 79 protein-coding and 4 rRNA genes from 20 plastomes as determined by RAxML (−ln L= −263,704.243992). The numbers at each node indicate the ML bootstrap values. Genbank accession numbers of taxa are shown in the following: Abelmoschus esculentus (NC_035234), Althaea officinalis (NC_034701), Aquilaria sinensis (NC_029243), Arabidopsis thaliana (NC_000932), Brassica napus (NC_016734), Citrus sinensis (NC_008334), Durio zibethinus (MG138151, this study), Gossypium arboreum (NC_016712), Gossypium barbadense (NC_008641), Gossypium robinsonii (NC_018113), Gossypium thurberi (NC_015204), Hibiscus syriacus (KR259989), Panax ginseng (NC_006290), Sesamum indicum (NC_016433), Talipariti hamabo (NC_030195), Theobroma cacao (NC_014676), Tilia amurensis (NC_028588), Tilia mandshurica (NC_028589), Tilia oliveri (NC_028590), and Tilia paucicostata (NC_028591).