Figures & data
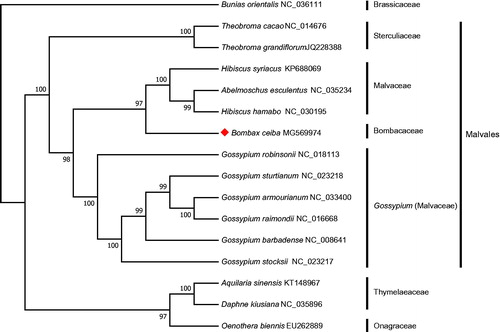
Figure 1. The maximum-likelihood phylogenetic tree constructed with chloroplast genomes of 16 plants. Bootstrap values are shown at the nodes. The chloroplast genome accession number used in the phylogeny study: Bunias orientalis: NC_036111; Theobroma cacao: NC_014676; Theobroma grandiflorum: JQ228388; Hibiscus syriacus: KP688069; Abelmoschus esculentus: NC_035234; Hibiscus hamabo: NC_030195; Bombax ceiba: MG569974; Gossypium robinsonii: NC_018113; Gossypium sturtianum: NC_023218; Gossypium armourianum: NC_033400; Gossypium raimondii: NC_016668; Gossypium barbadense: NC_008641; Gossypium stocksii: NC_023217; Aquilaria sinensis: KT148967; Daphne kiusiana: NC_035896; Oenothera biennis: EU262889.