Figures & data
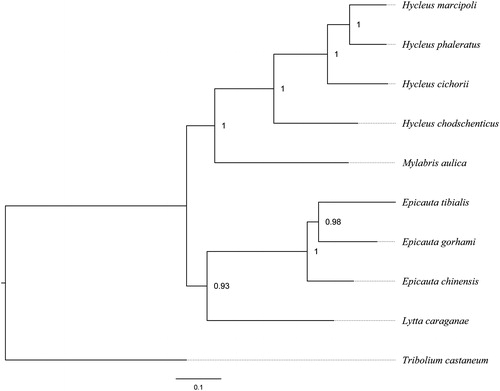
Figure 1. The Bayesian phylogenetic tree of nine meloids was constructed using the dataset of 13 protein-coding genes, with the tenebrionoid Tribolium castaneum employed as the outgroup. The numbers abutting nodes refer to Bayesian posterior probabilities. Sequence data used in this study are the following: Hycleus marcipoli (KX161857), Hycleus chodschenticus (KT808466), Mylabris aulica (KX161860), Epicauta chinensis (KP692789), Epicauta tibialis (KX161855), Epicauta gorhami (KX161854), Lytta caraganae (KX161859), Tribolium confusum (NC_026702).