Figures & data
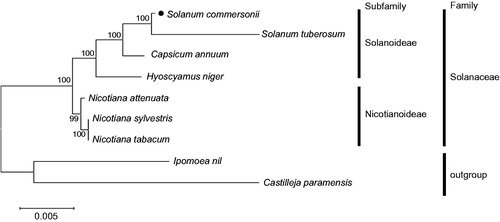
Figure 1. Maximum likelihood (ML) tree of nine Solanaceae-related species. The 23 protein-coding sequences in mitochondrial common gene were aligned using MAFFT (http://mafft.cbrc.jp/alignment/server/index.html). And then, phylogenetic tree was constructed with tamura-nei model and 1000 bootstrap method by MEGA 7.0 (Kumar et al. Citation2016). Mitochondrial genome sequences used for this tree are Capsicum annuum, NC_024624; Castilleja paramensis, NC_031806; Hyoscyamus niger, NC_026515; Ipomoea nil, NC_031158; Nicotiana attenuate, MF579563; N. sylvestris, NC_029805; N. tabacum, NC_006581; S. commersonii, MF989960 and MF989961; S. tuberosum, MF989953 to MF989957.