Figures & data
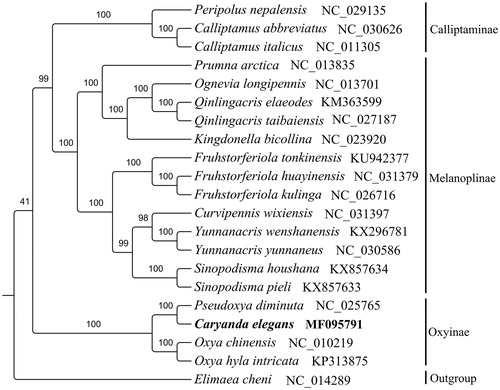
Figure 1. Maximum likelihood tree obtained using RaxML (Stamatakis Citation2014) with 1000 non-parametric bootstrap replicates. DNA sequences were aligned with MEGA4.0 (Tamura et al. Citation2007) individually and alignments of individual genes were concatenated using SequenceMatrix v1.7.8 (Vaidya et al. Citation2011). PartitionFinderV1.1.1 (Lanfear et al. Citation2012) was used to determine the optional model of evolution and partitioning scheme simultaneously. All mitogenomes were downloaded from GenBank except C. elegans, and the accession number was given with species names.