Figures & data
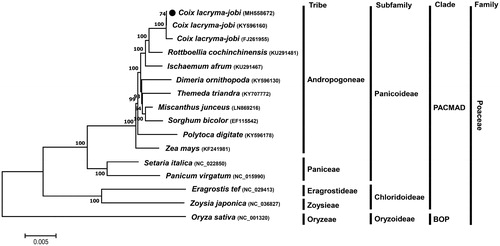
Figure 1. ML phylogenetic tree of chloroplast genomes of C. lacryma-jobi var.ma-yuen cv. Johyun and other Poaceae species. Whole chloroplast genome sequences were multiple-aligned using MAFFT (http://mafft.cbrc.jp/alignment/server/index.html) and used to generate phylogenetic tree by MEGA 7.0 (Kumar et al., Citation2016). The bootstrap support values (>50%) from 1000 replicates are indicated on the nodes. GenBank accession nos. of chloroplast genome sequences used for this tree are indicated within parentheses.