Figures & data
Figure 1. (A) Map showing the collection localities of F. occidentalis in Karnataka states of southern India with latitude and longitude. (B) Bayesian (BA) phylogeny of the studied F. occidentalis species. The BA posterior probability, and bootstrap support resulted in NJ and ML tree were superimposed with each nodes. The collection localities of the studied and database sequences were incorporated with the voucher IDs and accession numbers in the phylogeny. Colour bars show the genetic variability and distinct clades of the F. occidentalis population.
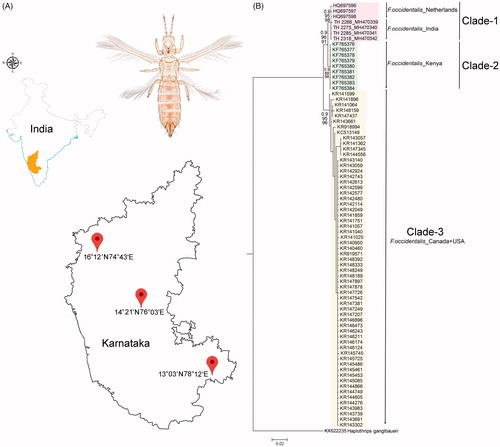
Table 1. The K2P genetic divergence over sequence pairs between and within groups of the studied F. occidentalis species.
