Figures & data
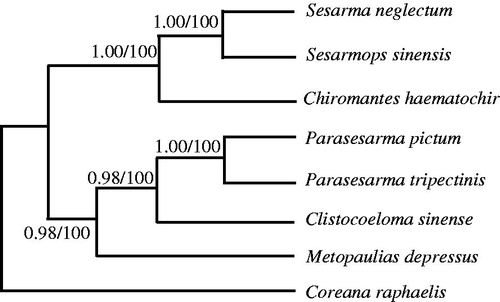
Figure 1. Phylogenetic relationships among the seven Sesarmidae species based on complete mtDNA sequences. Numbers at each node are Bayesian posterior probabilities (left) and maximum likelihood bootstrap proportions (estimated from 100 pseudoreplicates) (right). The accession number in GenBank of seven Sesarmidae in this study: Sesarmops sinensis (NC_031851), S. neglectum (NC_031851), Chiromantes haematochir (MH457175), Parasesarma tripectinis (NC_030046), P. pictum (MG580780), Metopaulias depressus (NC_030535), and Coreana raphaelis (NC_007976).