Figures & data
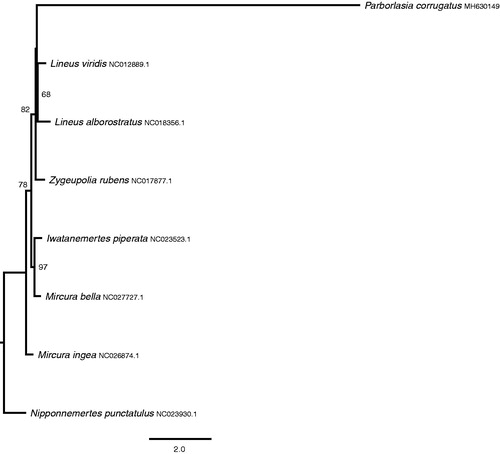
Figure 1. Maximum-likelihood tree with bootstrap support (1000 iterations) of concatenated amino acid sequences for 13 mitochondrial protein-coding genes in eight nemertean species (see text). Data were alignment in Mafft (Katoh and Standley Citation2013) and a PROTGAMMAAUTO model was employed in RAxML (Stamatakis Citation2014). Bootstrap values above 50% are shown on the tree. The hoplonemertean Nipponnemertes punctatula was used as the outgroup.