Figures & data
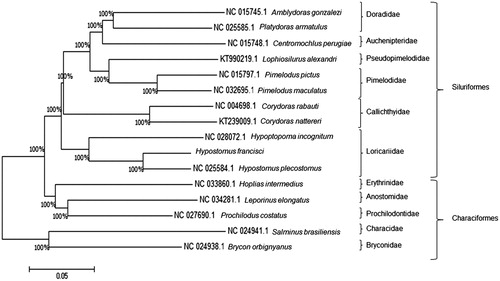
Figure 1. Tree of similarity of mitochondrial DNA (mtDNA) sequences. Comparison of the mitogenomes of ten species of the Order Siluriformes: Hypostomus Plecostomus (NC_025584.1), Hypoptopoma incognitum (NC_028072.1), Amblydoras gonzalezi (NC_015745.1), Platydoras armatulus (NC_025585.1), Centromochlus perugiae (NC_015748.1), Corydoras rabauti (NC_004698.1), Corydoras nattereri (KT239009.1), Pimelodus pictus (NC_015797.1), Pimelodus maculatus (NC_032695.1), and Lophiosilurus alexandri (KT990219.1); and five species of the Order Characiformes: Hoplias intermedius (NC_033860.1), Salminus brasiliensis (NC_024941.1), Leporinus elongatus (NC_034281.1), Brycon orgignyanus (NC_024938.1), and Prochilodus costatus (NC_027690.1). Characiform species were used as the outgroup. The consensus tree was constructed using the Kimura-2 parameter model (Kimura Citation1980) and 1000 bootstrap. The D-loop region was excluded from this analysis due to its high variability (Gonder et al. Citation2007). Hypostomus francisci grouped with H. plecostomus, and together they formed a sister group of H. incognitum, thus maintaining the Family Loricariidae as a single clade. Percentage support values for each group are indicated on the branches.