Figures & data
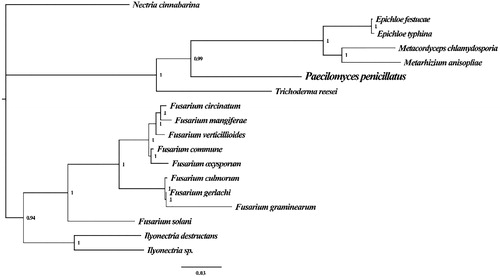
Figure 1. Molecular phylogenies of 18 species based on Bayesian inference analysis of the combined mitochondrial gene set (15 core protein-coding genes). Node support values are Bayesian posterior probabilities (BPP). Mitogenome accession numbers used in this phylogeny analysis: Epichloe festucae (NC_032064), Epichloe typhina (NC_032063), Metacordyceps chlamydosporia (NC_022835), Metarhizium anisopliae (NC_008068), Trichoderma reesei (NC_003388), Fusarium circinatum (NC_022681), Fusarium mangiferae (NC_029194), Fusarium verticillioides (NC_016687), Fusarium commune (NC_036106), Fusarium oxysporum (NC_017930), Fusarium culmorum (NC_026993), Fusarium gerlachii (NC_025928), Fusarium graminearum (NC_009493), Fusarium solani (NC_016680), Ilyonectria destructans (NC_030340), Ilyonectria sp. (MF924828), Nectria cinnabarina (NC_030252).