Figures & data
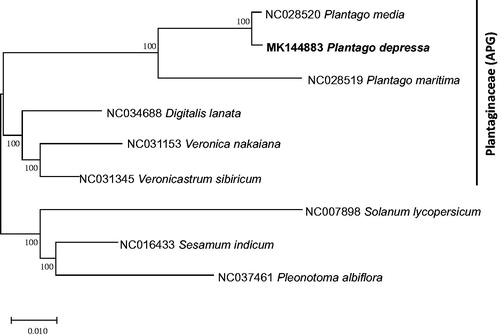
Figure 1. Neighbor joining and Maximum parsimony phylogenetic tree (bootstrap repeat is 10,000) of six Plantaginaceae chloroplast genomes: Plantago depressa (MK144883; this study), Plantago media (NC_028520), Plantago maritima (NC_028519), Veronica nakaiana (NC_031153), Veronicastrum sibiricum (NC_031345), Pleonotoma albiflora (NC_037461), Digitalis lanata (NC_034688), Sesamum indicum (NC_016433), and Solanum lycoverscum (NC_007898). The numbers above branches indicate bootstrap support value of neighbor joining tree.