Figures & data
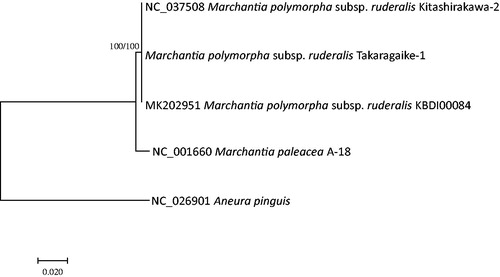
Figure 1. Maximum likelihood (bootstrap repeat is 1,000) and neighbor joining (bootstrap repeat is 10,000) phylogenetic trees of liverworts based on five complete mitochondrial genomes: M. polymorpha subsp. ruderalis (MK202951, NC_037508, and assembled sequence from Takaragaike-1 strain), M. paleacea (NC_001660), and Aneura pinguis (NC_026901). The numbers above branches indicate bootstrap support values of maximum likelihood and neighbor joining trees, respectively.