Figures & data
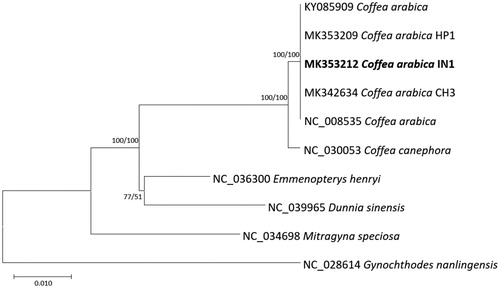
Figure 1. (A) Neighbor joining (bootstrap repeat is 10,000) and maximum likelihood (bootstrap repeat is 1,000) phylogenetic trees of five Coffea and four Rubiaceae complete chloroplast genomes: five Coffea arabica (MK353212, in this study, NC_008535, KY085909, MK342634, and MK353209), Coffea canephora (NC_030053), Mitragyna speciosa (NC_034698), Dunnia sinensis (NC_039965), Emmenopterys henryi (NC_036300), and Gynochthodes nanlingensis (NC_028614). Phylogenetic tree was drawn based on neighbor joining tree. The numbers above branches indicate bootstrap support values of neighbor joining and maximum likelihood phylogenetic trees, respectively.