Figures & data
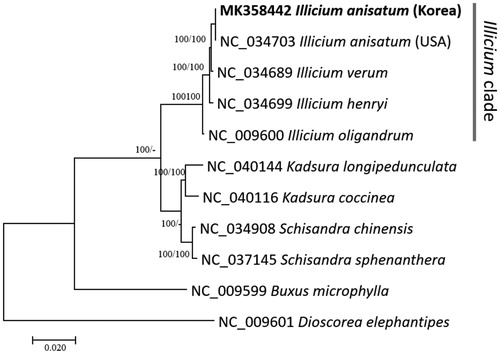
Figure 1. Neighborjoining (bootstrap repeat is 10,000) and maximum likelihood (bootstrap repeat is 1,000) phylogenetic trees of five Illicium and six Schisandraceae chloroplast genomes: Illicium anisatum (MK358442 in this study and NC_034703), Illicium verum (NC_034689), Illicium henryi (NC_034699), Illicium oligandrum (NC_009600), Kadsura longipedunculata (NC_040144), Kadsura coccinea (NC_040116), Schisandra chinensis (NC_034908), Schisandra sphenanthera (NC_037145), Buxus microphylla (NC_009599), and Dioscorea elephantipes (NC_009601). Phylogenetic tree was drawn based on neighbor joining tree. The numbers above branches indicate bootstrap support values of maximum likelihood and neighbor joining phylogenetic trees, respectively.