Figures & data
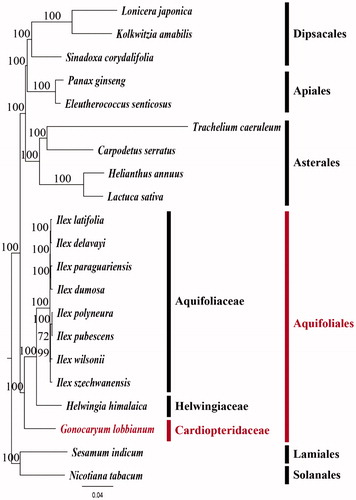
Figure 1. Maximum Likelihood (ML) tree based on 79 protein-coding and four rRNA genes from 21 plstomes as determined by RAxML(-ln L = 294507.154497). The numbers at each node indicate the ML bootstrap values. Genbank accession numbers of taxa are shown below, Calpodetus serratus (NC_036084), Eleutherococcus senticosus (NC_016430), Gonocaryum lobbianum (MK390345), Helianthus annuus (NC_007977), Helwingia himalaica (NC_031370), Ilex elavayi (KX426470), Ilex dumosa (KP016927), Ilex latifolia (KX426465), Ilex paraguariensis (KP016928), Ilex polyneura (KX426468), Ilex pubescens (KX426467), Ilex szechwanensis (KX426466), Ilex wilsonii (KX426471), Kolkwitzia amabilis (NC_029874), Lactuca sativa (NC_007578), Lonicera japonica (NC_026839), Nicotiana tabacum (NC_001879), Panax ginseng (NC_006290), Sesamum indicum (NC_016433), Sinadoxa corydalifolia (NC_032040), and Trachelium caeruleum (NC_010442).