Figures & data
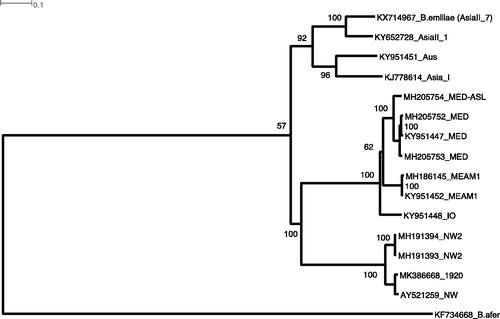
Figure 1. Maximum Likelihood phylogenetic analyses of 15 Bemisia tabaci cryptic species mitogenomes and one B. afer mitogenome as an outgroup using IQTree with 1000 bootstrap replications involving 10,556 bp representing 13 concatenated coding gene aligned sequences with end trimming and gaps moved. The historical 1920 Barbados B. tabaci ‘NW’ species (MK386668) clustered with the Sanger sequenced NW (AY521259, Thao et al. Citation2004), and together with the two NW2 mitogenomes (De Marchi et al. Citation2018), formed a ‘New World’ clade with 100% bootstrap value for node support. Note that the AsiaII_1 mitogenome (KY652728) was available as an unpublished genome from GenBank (accessed 07-Jan-2019).