Figures & data
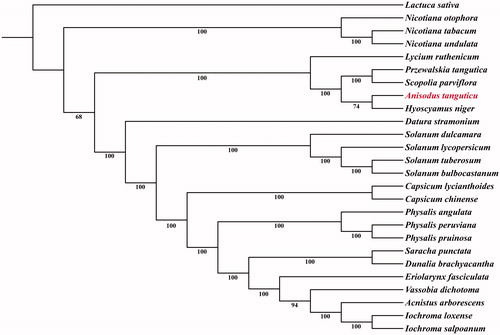
Figure 1. Maximum likelihood phylogenetic tree based on 26 complete chloroplast genome sequences. The number on each node indicates the bootstrap value. Accession numbers: Acnistus arborescens KU306735.1; Anisodus tanguticus MF539117; Capsicum chinense KU041709.1; Capsicum lycianthoides KP274856.1; Datura stramonium JN654342.1; Dunalia brachyacantha KP308151.1; Eriolarynx fasciculata KU306938.1; Hyoscyamus niger KF248009.1; Iochroma loxense KP296185.1; Iochroma salpoanum KU315119.1; Lactuca sativa AP007232; Lycium ruthenicum MG976805.1; Nicotiana otophora KU051626.1; Nicotiana tabacum Z00044.2; Nicotiana undulata JN563929.1; Physalis angulata MH019241.1; Physalis peruviana MH019242.1; Physalis pruinosa MH019243.1; Przewalskia tangutica KY352315.1; Saracha punctata KP280050.1; Solanum bulbocastanum DQ347958.1; Solanum dulcamara KY863443.1; Solanum lycopersicum HG975525.1; Scopolia parviflora KU900232.1; Solanum tuberosum DQ386163.2; Vassobia dichotoma KP294521.1.