Figures & data
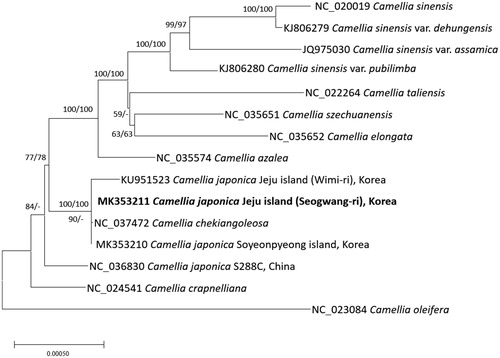
Figure 1. Neighbor joining (bootstrap repeat is 10,000) and maximum likelihood (bootstrap repeat is 1,000) phylogenetic trees of 15 Camellia chloroplast genomes: Camellia japonica (MK353211 in this study, MK353210, KU951523, and NC_036830), Camellila chekiangoleosa (NC_037472), Camellia crapnelliana (NC_024541), Camellia oleifera (NC_024541), Camellia azalea (NC_035574), Camellia sinensis (NC_024541), Camellia sienesis var. pubilimba (KJ806280), Camellia sienesis var. assamica (JQ975030), Camellia sienesis var. dehungensis (KJ806279), Camellia szechuanensis (NC_035651), Camellia tallensis (NC_022264), Camellia elongata (NC_035652), and Camellia oleifera (NC_023084). The numbers above branches indicate bootstrap support values of maximum likelihood and neighbor joining phylogenetic trees, respectively.