Figures & data
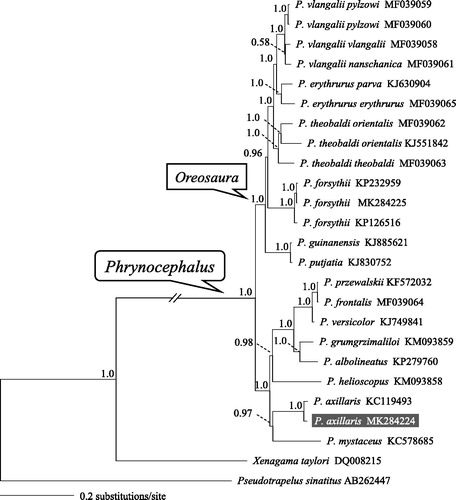
Figure 1. A majority-rule consensus tree inferred from Bayesian inference using MrBayes v.3.2.1 (Ronquist & Huelsenbeck Citation2003) with GTR + г substitution model, based on the concatenated PCGs of 22 individuals of toad-headed lizards and two outgroups. The novel sequencing sample is highlighted. GenBank accession numbers are given with species/subspecies names. DNA sequences were aligned in MEGA v.6.06 (Tamura et al. Citation2013). The PCGs were translated to amino acids sequences and were manually concatenated all sequences into a single nucleotide dataset (total 11,329 bp). Node numbers show Bayesian posterior probabilities. Branch lengths represent means of the posterior distribution.