Figures & data
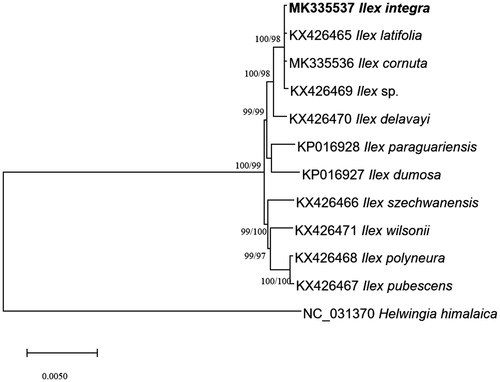
Figure 1. Neighbor joining (bootstrap repeat is 10,000) and maximum likelihood (bootstrap repeat is 1,000) phylogenetic trees of eleven Ilex and one Helwingia chloroplast genomes from Aquifoliaceae: Ilex integra (MK335537 in this study), Ilex cornuta (MK335536), Ilex latifolia (KX426465), Ilex sp. XY-2016 (KX426469), Ilex delavayi (KX426470), Ilex paraguaiensis (KP016928), Ilex dumorsa (KP016927), Ilex szechwanensis (KX426466), Ilex wilsonii (KX426471), Ilex polyneura (KX426468), Ilex pubescens (KX426467), and Helwingia himalaica (NC_031370). Section names were displayed in the right side of phylogenetic tree (Gottlieb et al. Citation2005; Wu et al. 2008; Jiang et al. Citation2017). Phylogenetic tree was drawn based on neighbor joining tree. The numbers above branches indicate bootstrap support values of maximum likelihood and neighbor joining phylogenetic trees, respectively.