Figures & data
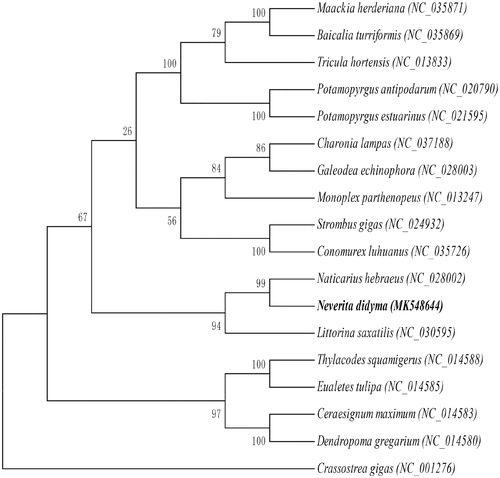
Figure 1. Maximum likelihood tree based on mitochondrial genome nucleotide sequences of the Neverita didyma (MK548644) and the other 17 species of Mollusca under the mutation model of GTR + I+G. Genbank accession numbers of the sequences were used for the tree as follows: Naticarius hebraeus (NC_028002.1), Charonia lampas (NC_037188.1), Baicalia turriformis (NC_035869.1), Maackia herderiana (NC_035871.1), Conomurex luhuanus (NC_035726.1), Galeodea echinophora (NC_028003.1), Ceraesignum maximum (NC_014583.1), Potamopyrgus estuarinus (NC_021595.1), Potamopyrgus antipodarum (NC_020790.1), Littorina saxatilis (NC_030595.1), Strombus gigas (NC_024932.1), Thylacodes squamigerus (NC_014588.1), Eualetes tulipa (NC_014585.1), Dendropoma gregarium (NC_014580.1), Tricula hortensis (NC_013833.1), Monoplex parthenopeus (NC_013247.1), Crassostrea gigas (NC_001276.1).