Figures & data
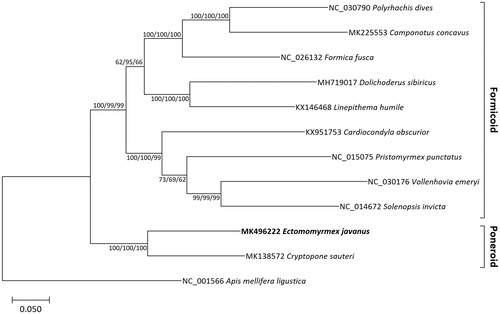
Figure 1. Maximum likelihood (bootstrap repeat is 1,000), neighbor-joining (bootstrap repeat is 10,000), and minimum evolution (bootstrap repeat is 10,000) phylogenetic trees of eleven ants and one honeybee mitochondrial genomes: Ectomomyrmex javanus (MK496222 in this study), Cryptopone sauteri (MK138572), Camponotus concavus (MK225553), Polyrhachis dives (NC_030790), Formica fusca (NC_026132), Solenopsis invicta (NC_014672), Cardiocondyla obscurior (KX951753), Vollenhovia emeryi (NC_030176), Pristomyrmex punctatus (NC_015075), Dolichoderus sibiricus (MH719017), Linepithema humile (KX146468), and Apis mellifera ligustica (NC_001566) as an outgroup. Phylogenetic tree was drawn based on maximum likelihood tree. The numbers above branches indicate bootstrap support values of maximum likelihood, neighbor joining, and minimum evolution phylogenetic trees, respectively.