Figures & data
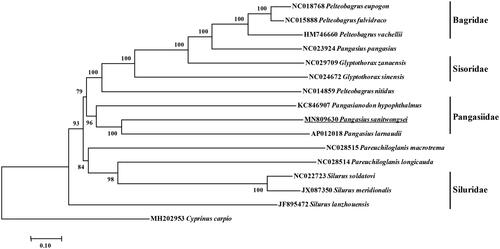
Figure 1. Molecular phylogeny of Pangasius sanitwongse and other Siluriformes varieties based on complete mitogenome. The phylogenic tree is constructed by neighbor joining method with 1000 bootstrap replicates. The mtDNA sequences are downloaded from Genbank, The sequence data for phylogenetic analyses used in this study were as follows: Pelteobagrus eupogon (NC018768), Pelteobagrus fulvidraco (NC015888), Pelteobagrus vachellii (HM746660), Pangasius pangasius (NC023924), Glyptothorax sinensis (NC024672), Glyptothorax zanaensis (NC029709), Pelteobagrus nitidus (NC014859), Pangasianodon hypophthalmus (KC846907), Pangasius larnaudii (AP012018), Pareuchiloglanis macrotrema (NC028515), Pareuchiloglanis longicauda (NC028514), Silurus soldatovi (NC022723), Silurus meridionalis (JX087350), Silurus lanzhouensis (JF895472) and Cyprinus carpio (MH202953).