Figures & data
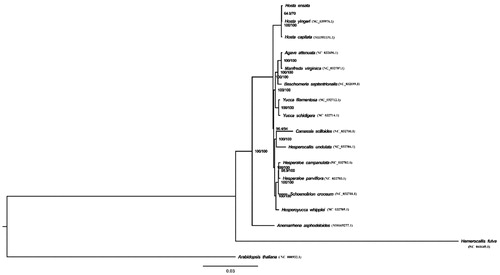
Figure 1. The phylogenetic tree was constructed using chloroplast genome sequences of 17 species within the Asparagaceae family and Arabidopsis thaliana as an outgroup based on the maximum-likelihood analysis using 500 bootstrap replicates. Chloroplast genome sequences used for this tree are Anemarrhena asphodeloides, NC_032698.1; Anemarrhena asphodeloides, MH669277.1; Arabidopsis thaliana, NC_000932.1; Beschorneria septentrionalis, NC_032699.1; Camassia scilloides, NC_032700.1; Hemerocallis fulva, NC_041649.1; Hesperaloe parviflora, NC_032703.1; Hesperaloe campanulata, NC_032702.1; Hesperocallis undulata, NC_032704.1; Hesperoyucca whipplei, NC_032705.1; Hosta capitata, MH581151.1; Hosta yingeri, NC_039976.1; Hemerocallis fulva, NC_041649.1; Manfreda virginica, NC_032707.1; Schoenolirion croceum, NC_032710.1, Yucca filamentosa, NC_032712.1; Yucca schidigera, NC_032714.1.