Figures & data
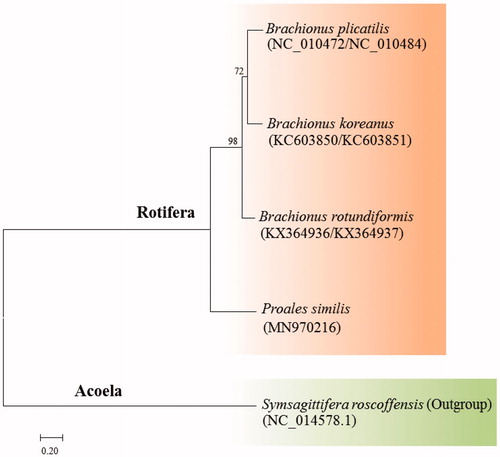
Figure 1. Phylogenetic analysis of the marine rotifer Proales similis mitochondrial DNA. We conducted a comparison of four monogonont rotifer species with 12 protein-coding genes (PCGs) with an outgroup. The amino acid sequences of 12 PCGs were aligned by ClustalW. Maximum likelihood (ML) analysis was performed by Raxml 8.2.8 (http://sco.h-its.org/exelixis/software.html) with GTR + γ+I nucleotide substitution model. The rapid bootstrap analysis was conducted with 1000 replications. The marine flatworm Symsagittifera roscoffensis served as an outgroup. Ln = −25855.214063.