Figures & data
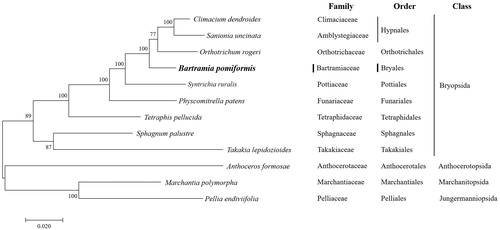
Figure 1. Phylogenetic position of B. pomiformis determined by Maximum-likelihood methods based on combined analysis with amino acids sequences of 16 mitochondrial genes common in all taxa. The bootstrap values (1000) are presented near the corresponding branch. Branches that were supported by above 50% bootstrap values are indicated by bold lines. Sequences from 2 Marchantiopsida and 1 Anthocerotophyta were used as outgroup. GenBank accession numbers of mitogenomes used are Anthoceros formosae (NC_004543), Bartramia pomiformis (MT024676), Climactium dendroides (MT006132), Marchantia polymorpha (NC_037507), Orthotrichum rogeri (NC_026212), Pellia endiviifolia (NC_019628), Physcomitrella patens (NC_037465), Sphagnum palustre (NC_03019), Syntrichia filaris (MK852705), Takakia lepidozioides (NC_028738) and Tetraphis pellucida (NC_024291).