Figures & data
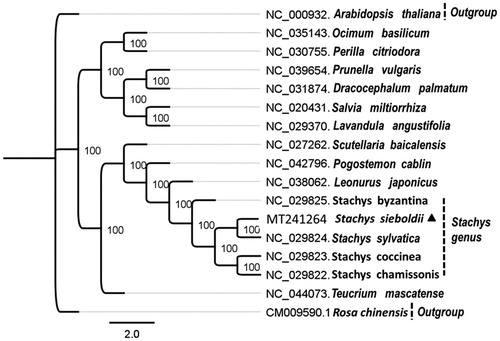
Figure 1. Maximum-likelihood phylogenetic tree base on 17 completely chloroplast genomes. The accession numbers are shown in the figure. Bootstrap support values based on 1000 replicates are displayed on each node. Rosa chinensis and Arabidopsis thaliana are used as outgroup. Marked by a black triangle is S. sieboldii in this study.