Figures & data
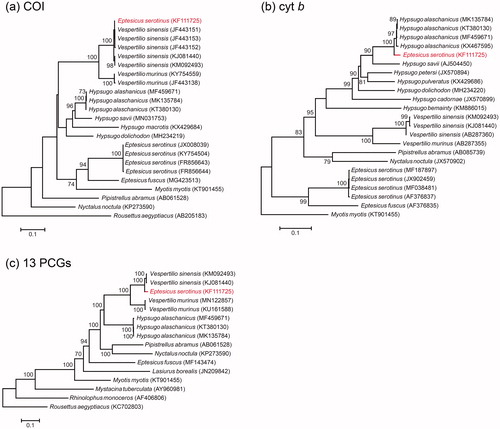
Figure 1. ML phylogenies of vespertilionid bats and selected outgroups based on (a) COI (698 bp), (b) cytochrome b (1140 bp) and (c) mitogenomes (13 PCGs, trimmed with GBLOCKS; 11,328 bp). Numbers along branches represent bootstrap support values (>70%) based on 1000 pseudoreplications. Note that KF111725 (Eptesicus serotinus) clustered with Vespertilio sinensis in the COI gene tree, but with Hypsugo alashanicus in the cytochrome b gene tree.
Data availability statement
The data that support the findings of this study are openly available on GenBank at https://www.ncbi.nlm.nih.gov/nucleotide. Accession numbers are listed in .
