Figures & data
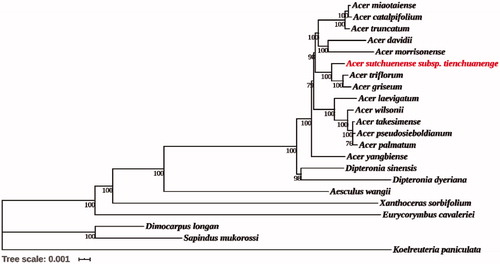
Figure 1. Phylogenetic tree reconstruction of 22 species using maximum likelihood (ML) based on the complete chloroplast genome sequences. There are the bootstrap support values from 1000 replicates given at each node. Their accession numbers are as follows: Acer catalpifolium: NC_041080; Acer davidii: NC_030331; Acer griseum: NC_034346; Acer pseudosieboldianum: NC_046487; Acer laevigatum: NC_042443; Acer miaotaiense: NC_030343; Acer morrisonense: NC_029371; Acer palmatum: NC_034932; Acer takesimense: NC_046488; Acer triflorum: NC_047296; Acer truncatum: NC_037211; Acer wilsonii: NC_040988; Acer yangbiense: MK479229; Aesculus wangii: NC_035955; Dimocarpus longan: NC_037447; Dipteronia dyeriana: NC_031899; Dipteronia sinensis: NC_029338; Eurycorymbus cavaleriei: NC_037443; Koelreuteria paniculata: NC_037176; Sapindus mukorossi: NC_025554; Xanthoceras sorbifolium: NC_037448.
Data availability statement
The data that support the findings of this study are openly available in GenBank of NCBI at https://www.ncbi.nlm.nih.gov/, reference number MT216764.
