Figures & data
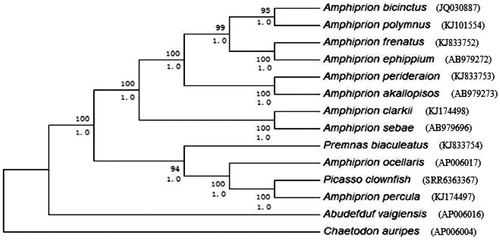
Figure 1. Phylogenetic trees derived from neighbour-joining (NJ) and Bayesian inference (BI) analyses based on nucleotide sequences of 13 mitochondrial protein-coding genes and 2 ribosomal RNA genes. Tree topologies produced by BI and NJ analyses were equivalent. Bayesian posterior probability (bottom) and bootstrap support values for NJ analyses (top) are shown in order on the nodes. Chaetodon auripes was selected as an outgroup species.
Data availability statement
The data that support the findings of this study are openly available in GenBank database at https://submit.ncbi.nlm.nih.gov/subs/sra/SUB3297187/overview, reference number [SRR6363367].
