Figures & data
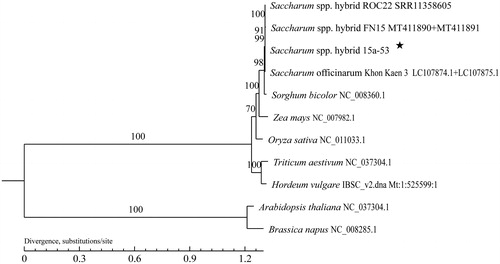
Figure 1. A maximum likelihood phylogenetic tree based on the comparison of mitochondrial genome sequences. GenBank accession numbers are listed after the species name. The numbers at the nodes are bootstrap percent probability values based on 1000 replications. The genome sequence in the present study is labeled with an asterisk.
Data availability statement
The data that support the findings of this study are available. The mitochondrial genome sequences rawdata of S. spp. hybrid 15a-53 were deposited in SRA with the accession number: SRR11358601 at the URL (https://www.ncbi.nlm.nih.gov/sra/?term=SRR11358601). The mitochondrial genome sequences of analyzed species (S. spp. hybrid FN15 and S. spp. hybrid ROC22; S. bicolor, Z. mays, O. sativa, T. aestivum, S. officinarum Khon Kaen 3, A. thaliana, and B. napus) were from the NCBI GenBank databases (the URL https://www.ncbi.nlm.nih.gov/genbank/) and the mitogenome of H. vulgare is available in Ensembl (http://asia.ensembl.org/index.html).
