Figures & data
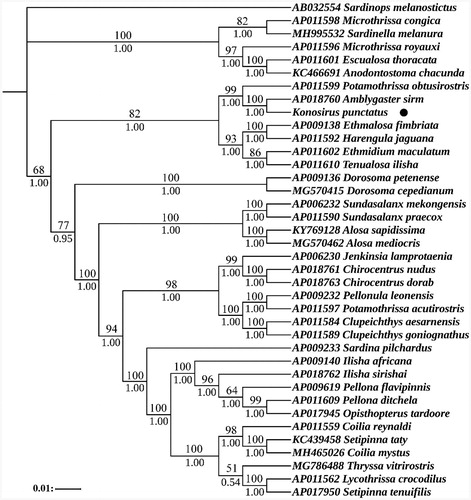
Figure 1. Phylogenetic analysis based on the nucleotide sequences of the 13 PCGs in the mitogenome. Support values for the Bayesian analyses (Bayesian posterior probabilities with 10 million generations; discarding 25% as burnin) and Maximum Likelihood analyses (bootstrap support with 1000 replications) are shown next to nodes. The number before the species name was the GenBank accession number. The numbers beside the nodes are posterior probabilities (BI, bottom) and bootstrap (ML, top). The genome sequence in this study was labeled with a black dot.
Data availability statement
The data that support the findings of this study are openly available in “NCBI” at https://www.ncbi.nlm.nih.gov/nuccore/MT801134.1, GenBank accession number: MT801134.1.
