Figures & data
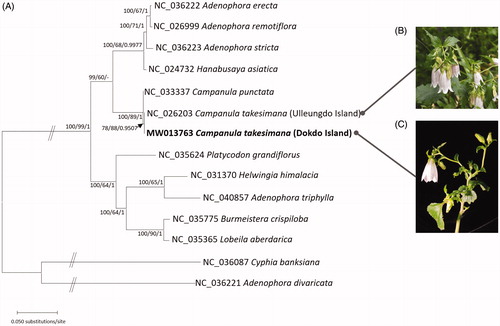
Figure 1. (A) Maximum-likelihood (bootstrap repeat is 1000) phylogenetic tree of 17 chloroplast genomes of Campanulaceae including two outgroups: Campanula takesimana (MW013763 in this study and NC_026203; Cheon et al. Citation2016), Campanula punctata (NC_033337; Yoo et al. Citation2016), Hanabusaya asiatica (NC_024732; Cheon and Yoo Citation2016), Adenophora stricta (NC_036223; Cheon et al. Citation2017), Adenophora erecta (NC_036222; Cheon et al. Citation2017), Adenophora remotiflora (NC_026999; Kim et al. Citation2016), Adenophora divaricata (NC_036221; Cheon et al. Citation2017), Adenophora triphylla (NC_040857), Platycodon grandifloras (NC_035624; Lin et al. Citation2019), Helwingia himalacia (NC_031370; Yao et al. Citation2016), Burmeistera crispiloba (NC_035775), Lobeila aberdarica (NC_035365), Cyphia banksiana (NC_036087). Neighbor-joining (bootstrap repeat is 10,000) and produced the same topology as the ML tree. The numbers above branches indicate bootstrap support values of maximum-likelihood, neighbor-joining, and Bayesian posterior probability, respectively. (B) Photograph of Campanula takesimana on Ulleungdo Island (taken by Yoonhyuk Bae). (C) Photograph of Campanula takesimana on Dokdo Island (taken by Hwa-Jung Suh).
Data availability statement
Mitochondrial genome sequence can be accessed via accession number MW013763 in GenBank of NCBI at https://www.ncbi.nlm.nih.gov. The associated BioProject, SRA, and Bio-Sample numbers are PRJNA668862, SAMN16425341, and SRR12816398, respectively.
