Figures & data
Figure 1. Comparison of the junction between inverted repeat region (IR), large single copy-region (LSC) and small single copy-region (SSC) of chloroplast genome among eight Coix species (varieties).
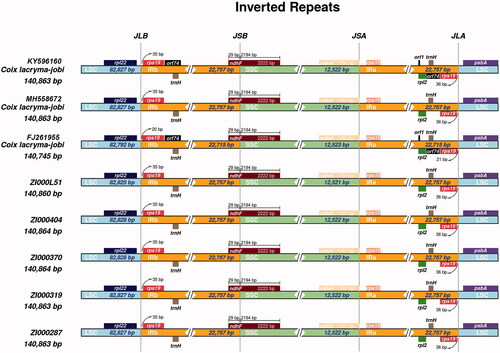
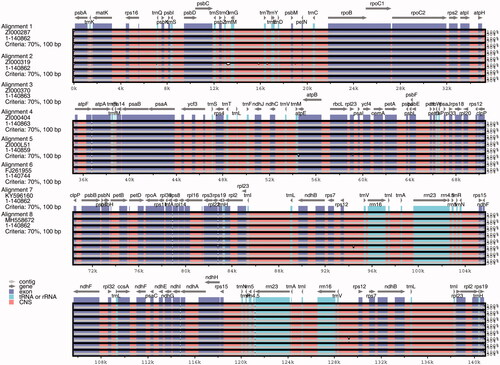
Figure 2. Comparison of eight chloroplast genomes of adlay. Note: Gray arrows and thick black lines above the alignment indicate gene orientation. Purple bars represent exons, pink bars represent conserved non-coding sequences (CNS), and blue bars represent mRNA. The y-axis represents the percentage identity (shown: 50–100%).
Data availability statement
The five genome sequences assembly has been deposited into NCBI GenBank under project ID: MT471102 (C. puellarum Balansa)(https://www.ncbi.nlm.nih.gov/nuccore/MT471102), MT471094 (C. stenocarpa Balansa)(https://www.ncbi.nlm.nih.gov/nuccore/MT471094), MT471095 (C.lacryma-jobi var. maxima Makino) (https://www.ncbi.nlm.nih.gov/nuccore/MT471095), MT471096 (C. chinensisvar. chinesis Tod.) (https://www.ncbi.nlm.nih.gov/nuccore/MT471096), MT471101 (C. chinensis var. formosana(Ohwi)L. Liu) (https://www.ncbi.nlm.nih.gov/nuccore/MT471101).