Figures & data
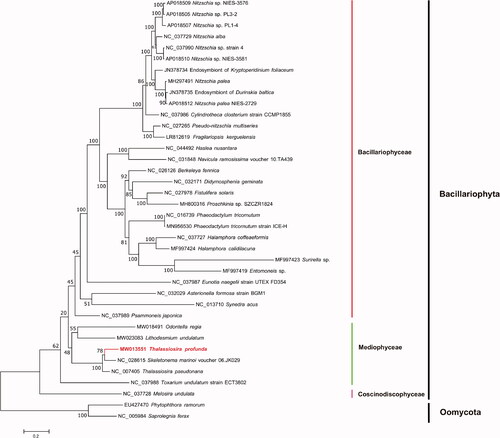
Figure 1. Maximum-likelihood (ML) phylogenetic tree based on tandem amino acid sequences of 31 common genes from 35 publicly diatom mitochondrial genomes, and Phytophthora ramorum (EU427470) and Saprolegnia ferax (NC_005984) in Oomycota were used as out-group taxa. The numbers beside branch nodes are the percentage of 1000 bootstrap values.
Data availability statement
The genome sequence data that support the findings of this study are openly available in GenBank of NCBI at https://www.ncbi.nlm.nih.gov/nuccore/MW013551, under the accession no. MW013551. The associated BioProject, SRA, and Bio-Sample numbers are PRJNA684688 (https://www.ncbi.nlm.nih.gov/bioproject/PRJNA684688), SRR13245496 (https://www.ncbi.nlm.nih.gov/sra/SRR13245496), and SAMN17065834 (https://www.ncbi.nlm.nih.gov/biosample/SAMN17065834/), respectively.
