Figures & data
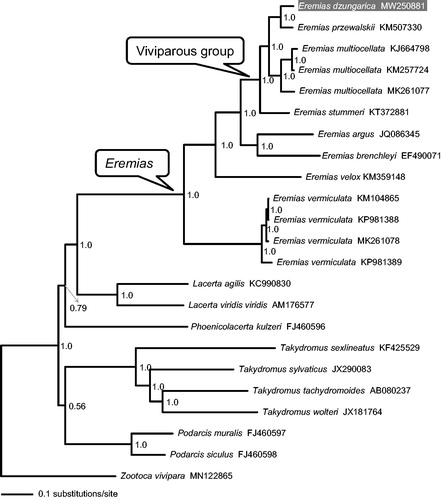
Figure 1. A majority-rule consensus tree inferred from Bayesian inference using MrBayes with the best models for each partition, based on the PCGs of Eremias spp. and ten other representative lacertids retrieved from GenBank. The phylogenetic placement of E. dzungarica is highlighted. GenBank accession numbers are given with species names. Node numbers show Bayesian posterior probabilities. Branch lengths represent means of the posterior distribution.
Data availability statement
The genome sequence data that support the findings of this study are openly available in GenBank of NCBI at (https://www.ncbi.nlm.nih.gov/nuccore/ MW250881) under the accession no. MW250881. The associated BioProject, SRA, and Bio-Sample numbers are PRJNA714298, SRR13960197, and SAMN18299467, respectively.
