Figures & data
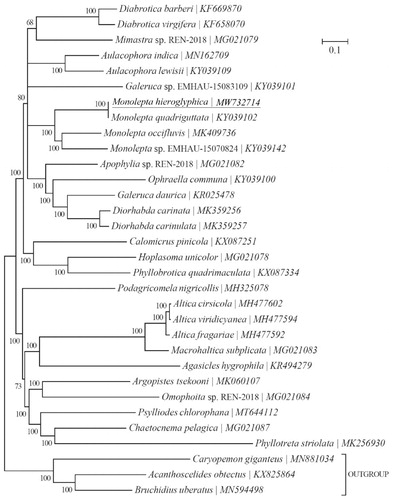
Figure 1. Phylogeny of twenty-nine Chrysomelidae-Galerucinae species based on 13 mitochondrial protein-coding genes reconstructed using Bayesian 3.2.0. The best-fit nucleotide substitution model is ‘GTR + G+I’. The support values are shown next to the nodes. Three Bruchinae species were included as outgroup taxa. Subfamily-level taxonomy was shown for each taxon.
Data availability statement
Mitochondrial genome sequence can be accessed via accession number MW732714 in GenBank of NCBI at https://www.ncbi.nlm.nih.gov/. Theassociated BioProject, SRA, and BioSample numbers are PRJNA713524, SRR13959601 and SAMN18253528, respectively.
