Figures & data
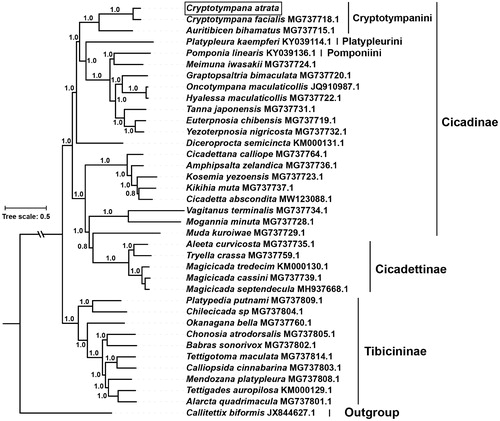
Figure 1. The phylogenetic tree inferred from Bayesian inference using MrBayes v.3.2.7 (Altekar et al. Citation2004) under GTR + I+G model, based on the concatenated 13 PCGs of 36 species of family Cicadidae and one outgroup. The novel sequencing mitogenome of C. atrata is highlighted with a box. GenBank accession numbers are given with species names. DNA sequences were aligned in muscle v3.8.31 (Edgar Citation2004). The numbers at branches indicate the Bayesian posterior probabilities. The corresponding generations of Bayesian inference when the standard deviation of split frequencies fell <0.01 was 41,000.
Data availability statement
The raw data for C. atrata has been deposited in the National Center for Biotechnology Information (NCBI database) SRA: BioProject ID PRJNA704537 at https://dataview.ncbi.nlm.nih.gov/object/SRR13775433. The assembled mitogenome of C. atrata in this study are openly available in the NCBI at https://www.ncbi.nlm.nih.gov/nucleotide/, reference number: MW405441.
