Figures & data
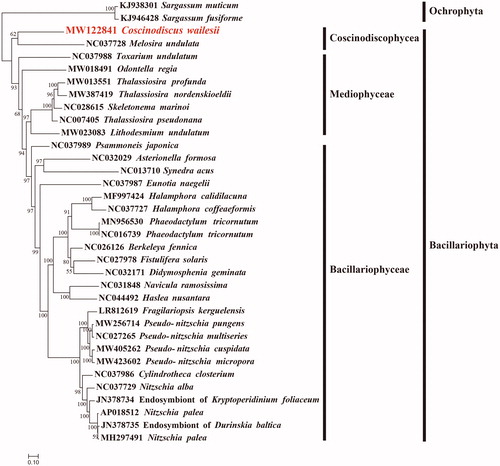
Figure 1. Maximum likelihood phylogenetic tree using concatenated amino acid sequences of 29 shared protein-coding genes (atp 8, 9; cob; cox1, 2, 3; nad1-7, 4 L, 9; rpl2, 5, 6, 14, 16; rps3, 4, 8, 10, 11, 13, 14, 19; and tatC) from 33 publicly available diatom mtDNAs, and Sargassum fusiforme (KJ946428) and Sargassum muticum (KJ938301) of Ochrophyta were used as out-group taxa. The numbers beside branch nodes are the percentage of 1000 bootstrap values.
Data availability statement
The genome sequence data that support the findings of this study are openly available in GenBank of NCBI at https://www.ncbi.nlm.nih.gov/under the accession number MW122841. The associated BioProject, SRA and Bio-Sample numbers are PRJNA686182, SRR13269809 and SAMN17108303, respectively.
