Figures & data
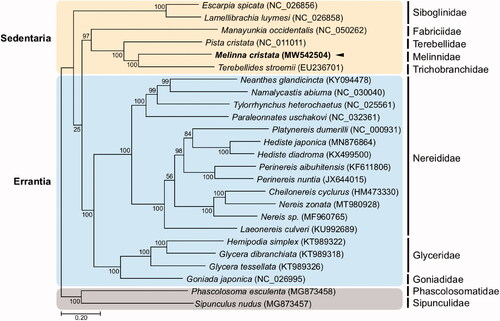
Figure 1. Maximum-likelihood (ML) phylogeny of 6 published mitogenomes from Sedentaria including M. cristata and 17 registered mitogenomes of Errantia species, and two Sipuncula species as an outgroup based on the concatenated nucleotide sequences of protein-coding genes (PCGs). Numbers on the branches indicate ML bootstrap percentages. DDBJ/EMBL/GenBank accession numbers for published sequences are incorporated. The black triangle means the polychaete analyzed in this study.
Data availability statement
BioProject, SRA, and BioSample accession numbers are https://www.ncbi.nlm.nih.gov/bioproject/PRJNA695143 and https://www.ncbi.nlm.nih.gov/biosample/SAMN17602389, respectively. The data that support the findings of this study are openly available in the National Center for Biotechnology Information (NCBI) at https://www.ncbi.nlm.nih.gov, with an accession number MW542504.
