Figures & data
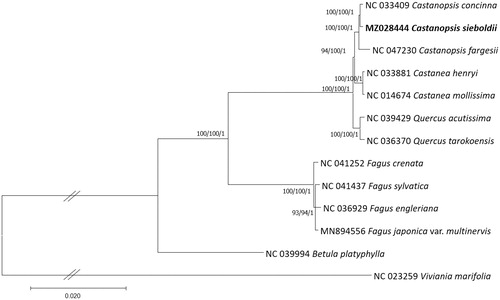
Figure 1. Maximum-Likelihood (bootstrap repeat is 1000), Neighbor-joining (bootstrap repeat is 10,000), and Bayesian inference (Number of generations is 1,100,000) phylogenetic trees of Fagaceae thirteen chloroplast genomes. Phylogenetic tree was drawn based on Maximum-Likelihood tree. The numbers above branches indicate bootstrap support values of Maximum-Likelihood, Neighbor-joining, and Bayesian inference phylogenetic trees, respectively.
Data availability statement
Chloroplast genome sequence can be accessed via accession number of MZ028444 in GenBank of NCBI at https://www.ncbi.nlm.nih.gov. The associated BioProject, SRA, and Bio-Sample numbers are PRJNA724746, SAMN18857600, and SRR14316975, respectively.
