Figures & data
Figure 1. The chloroplast genome of Phalaenopsis zhejiangensis. From the center going outward, the four circles indicate scattered forward and reverse repeats, tandem repeats, microsatellite sequences identified, and gene structure of the plastome.
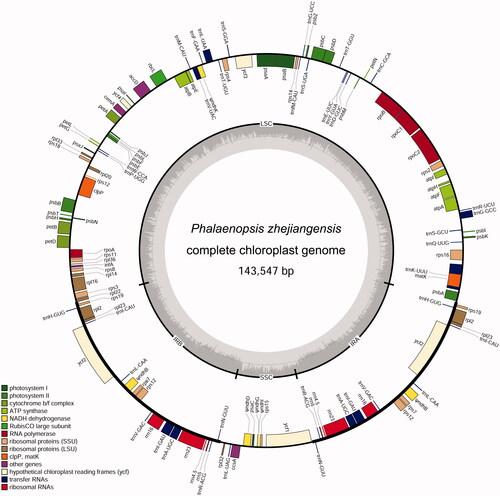
Table 1. List of genes in the chloroplast genome of Phalaenopsis zhejiangensis.
Figure 2. Structural variations at inverted-repeat and single-copy borders in nine Phalaenopsis chloroplast genomes. The figure features are not to scale regarding sequence length.
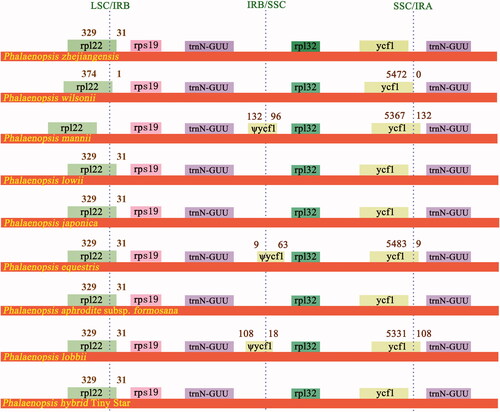
Table 2. Details of nine Phalaenopsis chloroplast genomes.
Table 3. Repeat sequences and their distribution within Phalaenopsis zhejiangensis chloroplast genome.
Data availability statement
The data that support the findings of this study are openly available in GenBank of NCBI at https://www.ncbi.nlm.nih.gov/nuccore/MZ326749. The associated BioProject, SRA, and Bio-Sample numbers are PRJNA734444, SRR14710950, and SAMN19491762, respectively.