Figures & data
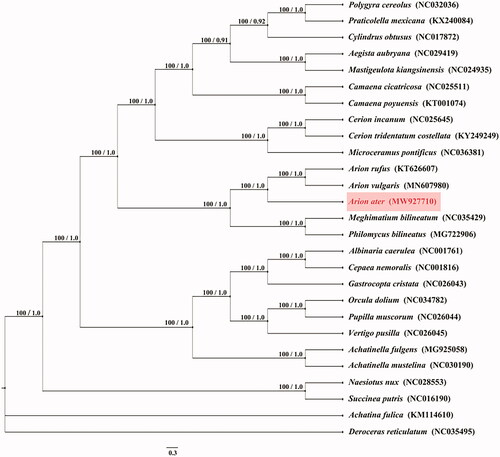
Figure 1. Phylogenetic tree of 27 species was obtain from maximum-likelihood (ML) and Bayesian phylogenetic inference (BI) method based on all protein-coding genes, the ML bootstrap proportions and BI posterior probabilities are shown on the nodes. Other mitochondrial genomes were downloaded from the NCBI Nucleotide databases (www.ncbi.nlm.nih.gov/nuccore).
Data availability statement
The genome sequence data that support the findings of this study are openly available in GenBank of NCBI (https://www.ncbi.nlm.nih.gov) under the accession no. MW927710. A total of sequences were downloaded from GenBank (http://www.ncbi.nlm.nih.gov/). The associated BioProject, SRA, and Bio-Sample numbers are PRJEB21599, ERS1807597, and SAMEA104148579, respectively.
