Figures & data
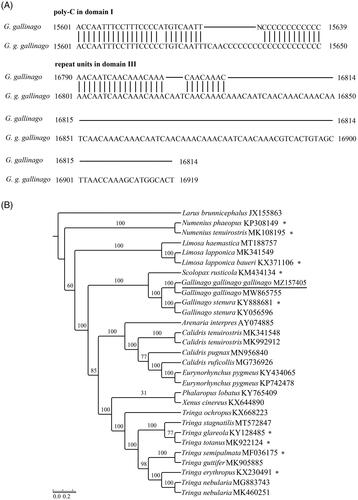
Figure 1. (A) Main nucleotide trait differences between G. gallinago (GenBank accession no. MW865755, MZ157405). (B) Maximum-likelihood tree obtained using RAxML v7.0.3 with 1000 nonparametric bootstrap replicates. GenBank accession numbers are indicated following species names. Numbers on the nodes are bootstrap values. ‘*’ represents no cytosine insertion.
Data availability statement
The genome sequence data that support the findings of this study are openly available in NCBI GenBank (https://www.ncbi.nlm.nih.gov/) under accession no. MZ157405. The associated BioProject, BioSample, and SRA numbers are PRJNA727369, SAMN19016979, and SRR14424283, respectively.
